What is QUINT?
Tip
Visit EBRAINS for more information about QUINT.
The QUINT workflow comprises a suite of software designed to support atlas-based quantification of labelled features in series of histological images from mouse or rat brain. All the software have graphical user interfaces (GUIs), no coding ability required. The QUINT workflow generates object counts and percentage coverage per atlas-region, in addition to point clouds for visualising the features in 3D atlas space. It currently supports quantification relative to the following atlases:
Allen Mouse Brain Atlas Common Coordinate Framework version 3 (2015 and 2017) (CCFv3)
Waxholm Atlas of the Sprague Dawley rat, version 2, 3 and 4 (WHS rat brain atlas).
QUINT involves several defined steps:
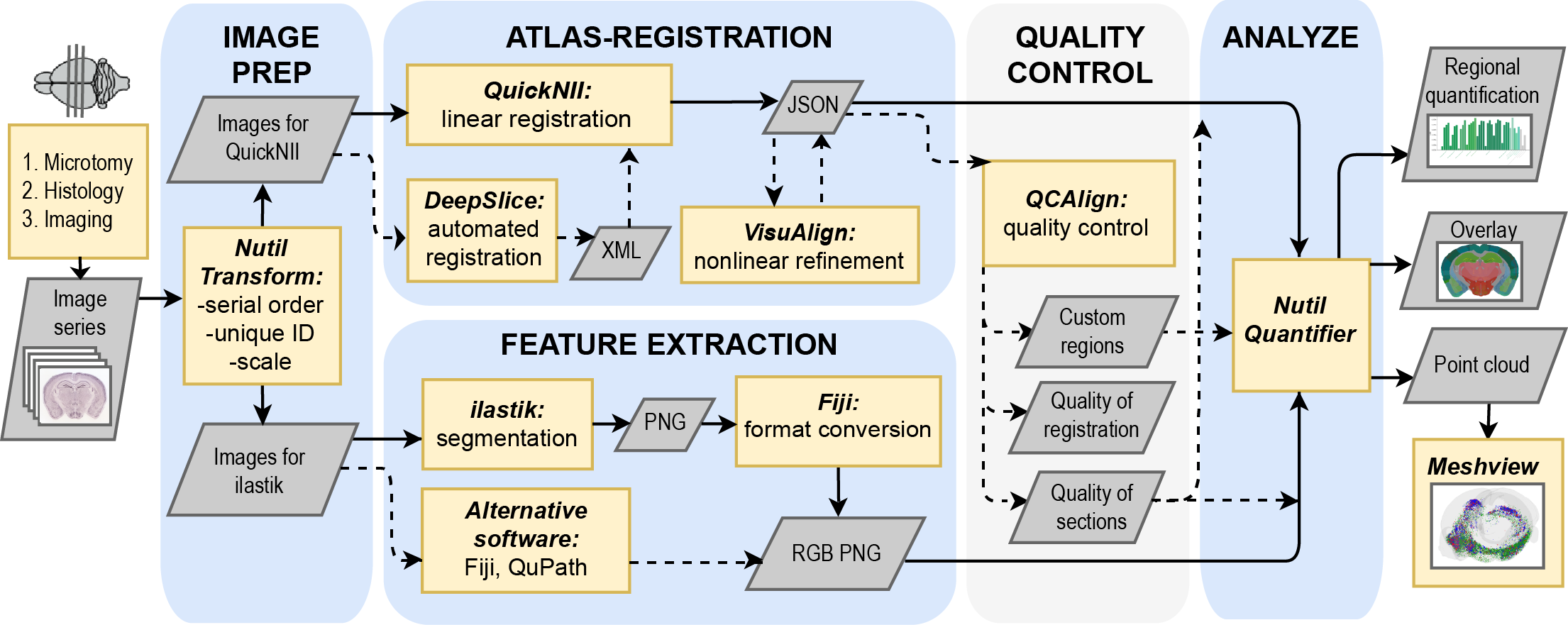
Image preparation using Nutil or another image analysis tool.
Atlas-registration with QuickNII, VisuAlign and/or DeepSlice. They support registration of the images to a reference brain atlas. Linear registration is performed using QuickNII, then VisuAlign enables refinement of the registration using non-linear methods. For coronal mouse brain sections, DeepSlice can automate the linear registration step.
Feature extraction using ilastik or another image analysis tool.
Quantification with Nutil.
Quality Control with QCAlign. This is optional and supports quality control of the section images, as well as quality control of the atlas-registration. It also enables exploration of the atlas hierarchy and creation of a customized hierarchy level to use for the quantification.
Visualisation in 3D with the Meshview Atlas Viewer.
Watch our video tutorial
Examples of use
QUINT supports histological sections from mouse and rat brain that have been labelled to reveal features. It works for sections cut at any angle of orientation, for complete as well as partial brain sections, and for sections affected by distortion. It can be used for:
Cell count analysis
To spatially characterise pathology
Analyse tract tracing connections
Check out the following articles that have used QUINT:
Lubben N et al. LRRK2 kinase inhibition reverses G2019S mutation-dependent effects on tau pathology progression. Translational Neurodegeneration. March 2024. http://doi.org/10.1186/s40035-024-00403-2
Geertsma et al. A topographical atlas of alpha-Synuclein dosage and cell-type expression in the adult mouse brain and peripheral organs. npj. March 2024. https://doi.org/10.1038/s41531-024-00672-8
Koen Seignette, Nora Jamann, Paolo Papale, Huub Terra, Ralph O Porneso, Leander de Kraker, Chris van der Togt, Maaike van der Aa, Paul Neering, Emma Ruimschotel, Pieter R Roelfsema, Jorrit S Montijn, Matthew W Self, Maarten HP Kole, Christiaan N Levelt. Jan 2024. Experience shapes chandelier cell function and structure in the visual cortex eLife 12:RP91153. https://doi.org/10.7554/eLife.91153.3
Guardamagna M, Chadney O, Stella F, Zhang Q, Kentros C, Battaglia F. Direct Entorhinal Control of CA1 Temporal Coding. bioRxiv. May 2023; doi: https://doi.org/10.1101/2023.05.27.542579
Goralski T et al. Spatial transcriptomics reveals molecular dysfunction associated with Lewy pathology. BioRxiv. May 2023. https://doi.org/10.1101/2023.05.17.541144
Zhang G, Xia S, Zhang N, Gao F, Zlokovik B, Zhang L, Zhao Z, Tao H. Integrative mapping of spatial transcriptomic and amyloid pathology in Alzheimer’s disease at single-cell resolution. Preprint. May 2023. https://doi.org/10.1101/2023.05.07.539389
Gurdon B, Yates SC, Csucs G, Groeneboom N, Hadad N, Telpoukhovskaia M, Oullette A, Oullette T, O’Connell K, Singh S, Murdy T, Merchant E, Bjerke I, Kleven H, Schlegel U, Leergaard T, Puchades M, Bjaalie J, Kaczorowski C. Detecting the effect of genetic diversity on brain composition in an Alzheimer’s disease mouse model. BioRxiv. Feb 2023. https://doi.org/10.1101/2023.02.27.530226
Hug NF, Lindsay NM, McCallum WM, Bryan J, Huang K, Ochadarena N, Tassou A, Scherrer G. Opioid receptor architecture for the modulation of brainstem functions. 2022 Dec. BioRxiv. https://doi.org/10.1101/2022.12.24.521865
Lopes MM, Paysan J, Rino J, Lopes SM, Pereira de Almeida L, Cortes L, Nobre RJ. A new protocol for whole-brain biodistribution analysis of AAVs by tissue clearing, light-sheet microscopy and semi-automated spatial quantification. Gene Ther. 2022 Nov 1. https://doi.org/10.1038/s41434-022-00372-z.
Jo Y, Lee SM, Jung T, Park G, Lee C, Im GH, Lee S, Park JS, Oh C, Kook G, Kim H, Kim S, Lee BC, Suh GSB, Kim SG, Kim J, Lee HJ. General-Purpose Ultrasound Neuromodulation System for Chronic, Closed-Loop Preclinical Studies in Freely Behaving Rodents. Adv Sci (Weinh). 2022 Oct 19:e2202345. https://doi.org/10.1002/advs.202202345
Botros P, Vendrell-Llopis N, Costa R, Carmena J. A neural model of proximity to reward. 2022 Oct. BioRxiv 2022.10.03.510669. https://doi.org/10.1101/2022.10.03.510669
Yao Y, Barger Z, Saffari Doost M, Tso CF, Darmohray D, Silverman D, Liu D, Ma C, Cetin A, Yao S, Zeng H, Dan Y. Cardiovascular baroreflex circuit moonlights in sleep control. Neuron. 2022 Sep 23:S0896-6273(22)00802-9. https://doi.org/10.1016/j.neuron.2022.08.027.
Ham GX, Augustine GJ. Topologically Organized Networks in the Claustrum Reflect Functional Modularization. Frontiers in Neuroanatomy. 16 June 2022. https://doi.org/10.3389/fnana.2022.901807
Bjerke IE, Cullity ER, Kjelsberg K, Charan KM, Leergaard TB, Kim JH. DOPAMAP, high-resolution images of dopamine 1 and 2 receptor expression in developing and adult mouse brains. Sci Data. 2022 Apr 19;9(1):175. https://doi.org/10.1038/s41597-022-01268-8
Telpoukhovskaia MA et al. Conserved cell-type specific signature of resilience to Alzheimer’s disease nominates role for excitatory cortical neurons. bioRxiv; doi: https://doi.org/10.1101/2022.04.12.487877
Tocco C, Øvsthus M, Bjaalie J.G, Leergaard T.B and Studer M. The topography of corticopontine projections is controlled by postmitotic expression of the area-mapping gene Nr2f1. Development; 149 (5). 2022. https://doi.org/10.1242/dev.200026
Kim S, Jo Y, Kook G, Pasquinelli C, Kim H, Kim K, Hoe HS, Choe Y, Rhim H, Thielscher A, Kim J, Lee HJ. Transcranial focused ultrasound stimulation with high spatial resolution. Brain Stimul. 2021 Mar-Apr;14(2):290-300. https://doi.org/10.1016/j.brs.2021.01.002
Whilden CM, Chevée M, An Seong Yeol, Pezon Brown S. The synaptic inputs and thalamic projections of two classes of layer 6 corticothalamic neurons in primary somatosensory cortex of the mouse. J Comp Neurol. 2021 Dec;529(17):3751-3771. doi: https://doi.org/10.1002/cne.25163. Epub 2021 May 6.
McDonald MW, Jeffers MS, Filadelfi M, Vicencio A, Heidenreich G, Wu J and Silasi G. Localizing Microemboli within the Rodent Brain through Block-Face Imaging and Atlas Registration. eNeuro 16 July 2021, 8 (4) ENEURO.0216-21.2021; DOI: https://doi.org/10.1523/ENEURO.0216-21.2021
Bjerke IE, Yates SC, Laja A, Witter MP, Puchades MA, Bjaalie JG and Leergaard TB. Densities and numbers of calbindin and parvalbumin positive neurons across the rat and mouse brain. 2021, iScience.https://doi.org/10.1016/j.isci.2020.101906